GANGE: Achieving Sequencing Without Sequencing With Diffusion Guided Generative Genomic Transformer
GANGE: Achieving Sequencing Without Sequencing With Diffusion Guided Generative Genomic Transformer
Gupta, S.; Kumar, A.; Bhati, U.; Shankar, R.
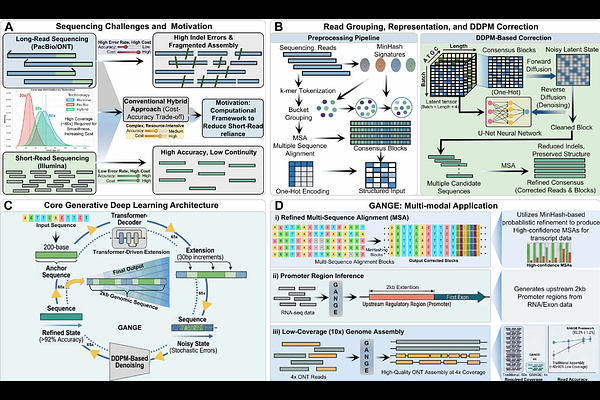
AbstractThe genome of a species is its book of life, but opening that book remains a costly affair due to the limitations the existing sequencing technologies pose. Short reads sequencers struggle to capture long and complex genomes, though have high fidelity rate. To counter that long reads from IIIrd generation sequencers are used, which are full of indel errors. Thus, reads from both approaches are collectively used with very high coverage, making the sequencing projects unreasonably high of cost and unapproachable to majority. Here we present a first of its kind generative deep-learning system, GANGE, which not just recovers the correct sequence with high accuracy from indel prone ONT reads at manifold lesser coverage but also extends it by 4kb, achieving sequencing without sequencing, horizontally as well as vertically while maintaining >92% accuracy consistently. This all makes it possible to drastically pull down sequencing project cost. GANGE was tested across A. thaliana, O. sativa genomes and Human chromosome 1 where it delivered outstanding assembly performance. Besides this, it was also used to accurately generate 2kb upstream promoters of all the genes from 12 different species, demonstrating that one can now also take up regulomics research just using RNA data alone when genome sequence is not available. With this all, GANGE brings a democratic turning point in the area of genomics and sequencing research.