Host background shapes the portability of a non-canonical translation initiation system across Escherichia coli strains
Host background shapes the portability of a non-canonical translation initiation system across Escherichia coli strains
Scopelliti, D.; Hutvagner, A.; Jaschke, P. R.
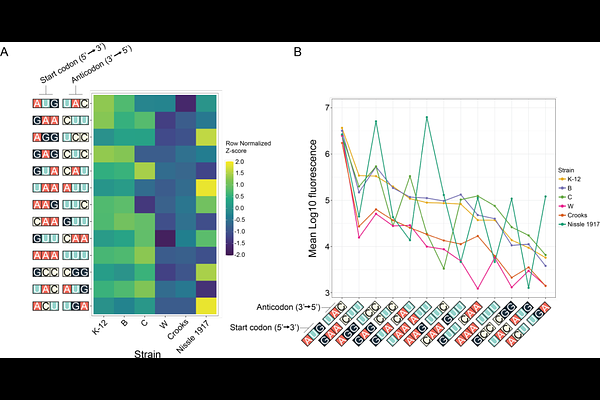
AbstractTranslation initiation has become an attractive target for engineering orthogonal translation systems, yet the extent to which these systems retain functionality across distinct host backgrounds remains poorly defined. In bacteria, start codon recognition depends on pairing between the initiator tRNA anticodon and a suitable start codon within the appropriate distance from the Shine-Dalgarno sequence. These sequence-specific interactions enable translation initiation to be reprogrammed through anticodon engineering. What is currently missing is an understanding of how anticodon mutants of initiator tRNAs function across different bacterial strains. Here, we systematically evaluated the portability of a library of twelve i-tRNA anticodon mutants paired with their complementary non-canonical start codons. Most i-tRNA-start codon pairs supported detectable translation initiation across multiple strains, demonstrating broad functional portability. However, initiation efficiency, absolute system output, and fitness effects varied substantially between strains. Comparative genomic analyses revealed host-specific gene differences broadly, and endogenous tRNA gene sequence and copy number specifically, was associated with this variability. While most i-tRNA variants were well tolerated, a subset produced strain-dependent growth defects that primarily affected growth rate rather than final culture density. Together, these findings show that translation initiation efficacy of engineered i-tRNAs is partially strain-dependent and that host background must be considered a key design variable when deploying these translation systems. Looking forward, this study provides a framework for host-aware selection of microbial chassis for orthogonal translation applications in synthetic biology.