Inferring division-associated stochasticity from time-series single-cell transcriptomes
Inferring division-associated stochasticity from time-series single-cell transcriptomes
Okochi, Y.; Sawazaki, Y.; Kondo, Y.; Naoki, H.
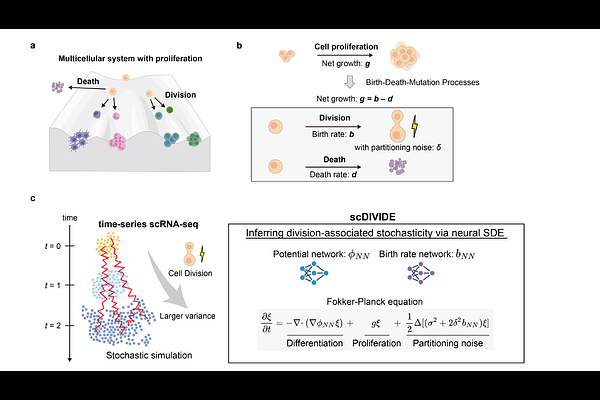
AbstractCell division is fundamental to multicellular organisms and stochastic partitioning of cellular components can strongly affect genome-wide gene expression states. However, how cell division-associated partitioning noise shapes the dynamics of proliferating cells is poorly understood. Here, we propose scDIVIDE, a neural stochastic differential equation framework to infer continuous cellular dynamics and division rates while accounting for partitioning noise. We combined birth-death-mutation processes from population genetics with dynamical optimal transport and revealed that the birth rate is embedded in the diffusion coefficient, enabling its inference from time-series scRNA-seq data. scDIVIDE accurately inferred birth rates in synthetic data and the inferred birth rates recapitulated turnover-related programs in mouse hematopoiesis data. By exploiting the birth-diffusion coupling, scDIVIDE provides a biologically-informed constraint on growth rate estimation, outperforming existing methods in predicting future cell distributions. scDIVIDE provides a conceptual avenue for quantitatively dissecting how partitioning noise shapes fate decisions in multicellular systems.