vcfilt: A Zero-Allocation Streaming Filter for High-Throughput VCF Processing
vcfilt: A Zero-Allocation Streaming Filter for High-Throughput VCF Processing
KP, M. M.
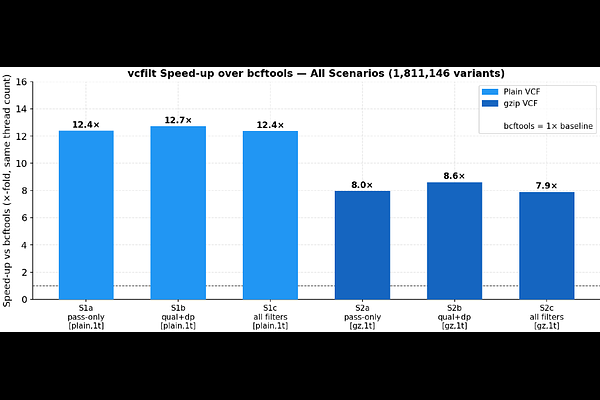
AbstractVariant Call Format (VCF) files are the dominant interchange format for genomic variant data, but their size - routinely exceeding tens of gigabytes for population-scale studies - creates a significant computational bottleneck at the quality-filtering stage. Existing tools such as bcftools and vcftools provide broad functionality through general-purpose expression engines, but incur substantial per-record overhead from dynamic field lookup, type resolution, and heap allocation. We present vcfilt, a streaming, batch-parallel VCF filter implemented in Go that restricts its scope to three high-frequency filter criteria (INFO/DP, INFO/AF, and QUAL) and applies them via a zero-allocation byte-scan parser. Benchmarked on real 1000 Genomes Project data (chromosome 20, 1,811,146 variants), vcfilt achieves 147,000 variants/second on an 18 GB plain-text VCF file using a single thread - a 12.2x speedup over bcftools 1.18 under identical conditions. On gzip-compressed input, the speedup is 7.9x. Output is byte-for-byte identical to bcftools across all tested filter combinations. vcfilt is distributed as a self-contained static binary, a Docker image, and a Singularity-compatible container. The source code and all benchmark scripts are openly available under the MIT licence.