Ultra-sensitive FLORA-seq links cell-type-specific tRNAome dynamics to differentiation trajectories guiding therapeutic suppressor tRNA candidate selection
Ultra-sensitive FLORA-seq links cell-type-specific tRNAome dynamics to differentiation trajectories guiding therapeutic suppressor tRNA candidate selection
Li, H.; Zhou, Y.-J.; Zhang, Z.; Ge, J.-Y.; Zhou, Z.-C.; Zhu, W.-Y.; Wu, X.-Y.; Wu, J.; Tian, P.-Y.; Gao, Y.; Sun, J.; Liu, R.-J.
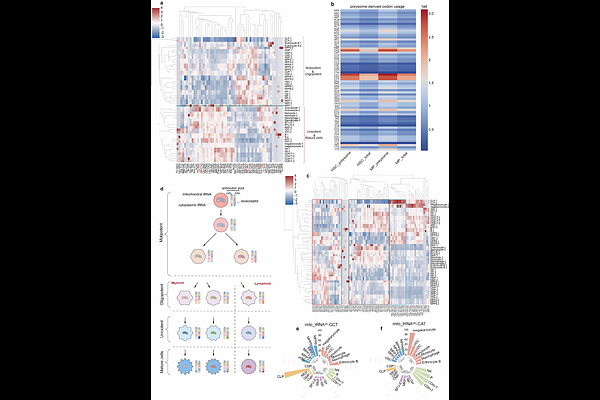
AbstractMammalian genomes encode hundreds of tRNA genes, but the role of individual tRNAs in development and cell identity remains unclear. Here, we introduce FLORA-seq, a method for simultaneous, low-input profiling of tRNAs, tRNA-derived RNAs (tdRs) and their modifications from as few as 5-20 cells. Applying FLORA-seq to mouse hematopoiesis revealed 50 high-variance isodecoders whose expression profiles recapitulate known hematopoietic cell types and correlate with differentiation trajectories. At the isoacceptor level, tRNA pools are largely stable within hematopoietic stem and progenitor cells (HSPCs), showing only subtle, specific variations, but become both distinct from HSPCs and internally stable within each terminally differentiated lineage. We further detected dynamic changes in tdR ratios and modification patterns, underscoring precise tRNA regulation during differentiation. Crucially, analysis of 54 endogenous tRNA isodecoders and their engineered suppressor counterparts showed that efficient premature termination codon readthrough preferentially arises from highly expressed cognate isodecoders. This correlation provides a rational framework for prioritizing suppressor tRNA candidates and implies non-redundant functions for individual isodecoders.