Benchmarking MSA pairing for protein-protein complex structure prediction reveals a depth-over-pairing principle
Benchmarking MSA pairing for protein-protein complex structure prediction reveals a depth-over-pairing principle
Luo, Y.; Wang, W.; Peng, Z.; Yang, J.
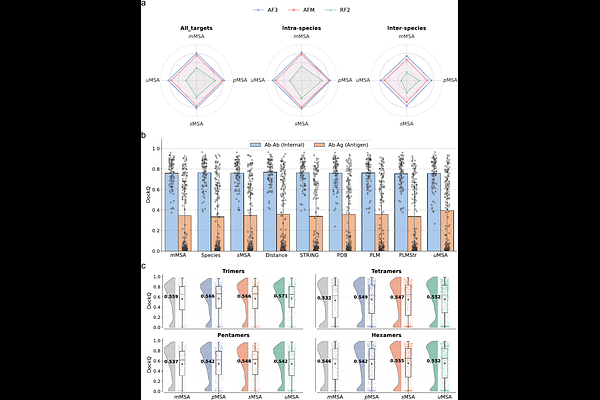
AbstractAlphaFold-Multimer (AFM) and AlphaFold3 (AF3) have revolutionized protein-protein complex structure prediction. However, it remains unclear whether elaborate MSA pairing is a strict prerequisite for achieving high accuracy. Here, we systematically evaluated diverse MSA construction strategies using a rigorous benchmark of 439 heterodimers. We found that incorporating paired MSAs yields only marginal improvements in structure prediction. Crucially, experiments using shuffled pairing patterns reveal that these gains stem not from specific pairing constraints, but rather from the increased depth of input MSAs, a finding that also extends to higher-order multimers. Furthermore, we demonstrate that prioritizing the inclusion of more homologs without pairing proves to be a superior strategy, enhancing performance for both intra- and inter-species dimers as well as the more challenging antibody-antigen complexes. This observation is substantiated by the recent success of the omicMSA strategy. Mechanistic analysis indicates that this capability is driven by inter-subunit physicochemical complementarity and the intrinsic architecture of the AF3 network. Furthermore, we identify critical bottlenecks that limit prediction accuracy, including large complex dimensions, restricted interface areas, and low experimental resolution. These findings establish a "depth-over-pairing" principle, offering new insights for improving protein complex prediction.