The RNase and RNA binding activities of selected RNase R truncations and mutations plus a detailed step by step protocol to purify recombinant RNase R
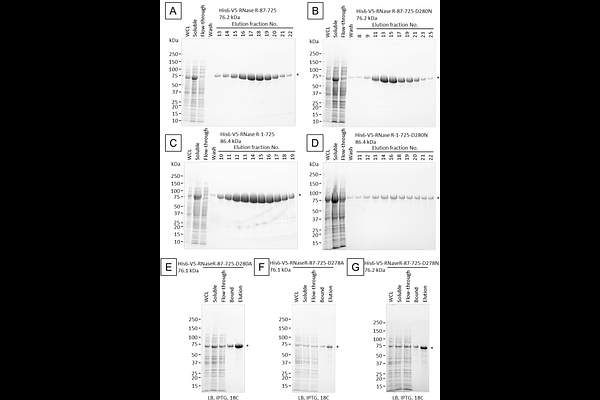
The RNase and RNA binding activities of selected RNase R truncations and mutations plus a detailed step by step protocol to purify recombinant RNase R
Horikawa, W.; Kiss, D. L.
AbstractRNase R is a processive 3' to 5' exoribonuclease that degrades a broad array of linear RNA species while preserving RNA lariats and circular RNAs (circRNA). In recent years, this enzyme has become pivotal for the field of circRNA research, serving as a key step for circRNA enrichment, purification, and identification. Despite this growing importance, the effects of mutations and truncations in RNase R have been incompletely studied. We make several point mutations and assay their effects on the ability of RNase R to bind and/or degrade RNA substrates. Our data show that selected active site mutations have varying effects on RNA binding and degradation. Furthermore, the increasing interest in circRNA-based RNA therapeutic platforms highlights an urgent need for RNase R in RNA molecular biology labs. However, the substantial cost of commercial RNase R remains a bottleneck, particularly for large-scale studies or the development of circRNA-based technologies. In this protocol, we offer a solution to that problem, namely a more accessible and cost-effective means of purifying high-quality and low-cost RNase R. We provide a highly detailed yet simplified, high-yield protocol that produces recombinant RNase R from Escherichia coli. The method uses a single-step Ni-NTA affinity chromatography procedure without proteolytic tag removal and is optimized for entry-level FPLC systems such as the AKTA Start, ensuring that high-purity enzyme production does not require specialized, high-end instrumentation. A second key feature is the establishment of an optimized reaction framework, including specific buffer compositions and defined enzyme-to-substrate ratios for the purified RNase R. The protocol achieves functional equivalence to premium commercial RNase R, ensuring complete linear RNA digestion without compromising the integrity of circRNA. The combination of a simplified purification workflow and a robust reaction protocol provides an accessible, cost-effective, and reliable solution for any molecular biology laboratory requiring high volumes of RNase R.