Sampling antibody conformational ensembles withABodyBuilder4-STEROIDS
Sampling antibody conformational ensembles withABodyBuilder4-STEROIDS
Spoendlin, F. C.; Cagiada, M.; Ifashe, K.; Vavourakis, O.; Deane, C. M.
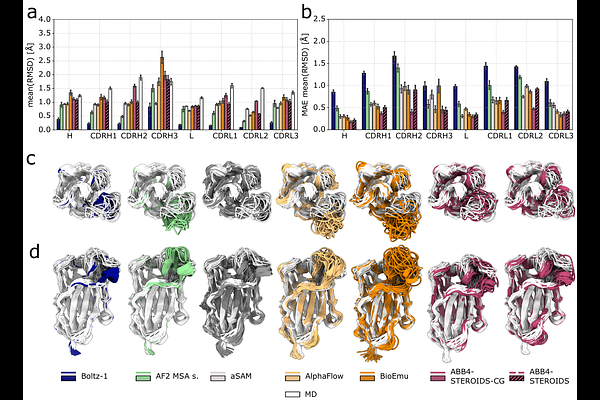
AbstractConformational flexibility is fundamental to the function of many proteins and in the case of antibodies can impact key properties such as affinity and specificity. While it is possible to predict single, static protein structures with high accuracy, predicting conformational ensemble remains challenging. Molecular dynamics simulations suffer from high computational costs, while deep learning methods are yet to achieve the same level of accuracy. Here, we introduce ABB4-STEROIDS a generative structure prediction model that samples conformational ensembles of antibodies. We trained our model on 4.2 million structural frames derived from $\sim$136,000 coarse-grained and a set of 83 new all-atom antibody MD simulations. We benchmarked our model on reproducing MD ensembles and evaluated the diversity of sampled structures and the covered conformational space against experimental evidence. ABB4-STEROIDS achieves state-of-the-art accuracy, particularly within the experimental benchmarks. The model is openly available and provides a robust resource for large-scale investigations of antibody conformational ensembles.