Evaluating Few-Shot Meta-Learning using STUNT for Microbiome-Based Disease Classification
Evaluating Few-Shot Meta-Learning using STUNT for Microbiome-Based Disease Classification
Peng, C.; Abeel, T.
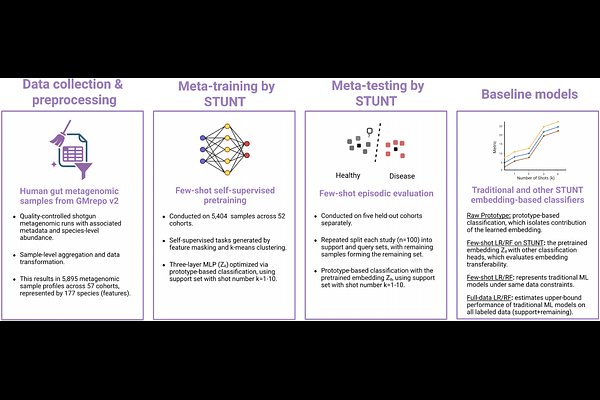
AbstractThe human gut microbiome is increasingly explored as a diagnostic indicator for disease, yet machine learning models trained on metagenomic data are often constrained by limited sample sizes and poor cross-cohort generalizability. Meta-learning, a machine learning paradigm that optimizes models for rapid adaptation to new tasks with limited examples, offers a promising strategy to address this by leveraging the potential shared microbial structure across publicly available metagenomic datasets. Here, we evaluated STUNT, a framework combining self-supervised pretraining with metric-based meta-learning (Prototypical Networks), for few-shot microbiome-based disease classification. Using over 5,000 species-level gut metagenomic profiles from 57 cohorts in GMrepo v2, we meta-trained STUNT on 52 cohorts and evaluated the pretrained embedding on five held-out disease cohorts covering rheumatoid arthritis (RA), gestational diabetes mellitus during pregnancy (GDM), non-alcoholic fatty liver disease (NAFLD), diabetes mellitus, type 1 (T1D), and inflammatory bowel disease (IBD). We compared Prototypical Networks, Logistic Regression, and Random Forest with and without STUNT-derived embeddings across shot sizes of 1 to 10 samples per class. We found that STUNT-derived embeddings provided a modest benefit only under extreme data scarcity (one labeled sample per class) and this advantage rapidly diminished and reversed with additional samples, indicating that the meta-learned representations impose an information bottleneck limiting access to task-specific signals. Classification performance varied substantially across cohorts, consistent with PERMANOVA-estimated microbiome-disease separability. These results highlight the need for representation learning approaches that preserve disease- and cohort-specific variation and suggest that intrinsic biological signal strength is the primary determinant of classification success.