cellNexus: Quality control, annotation, aggregation and analytical layers for the Human Cell Atlas data
cellNexus: Quality control, annotation, aggregation and analytical layers for the Human Cell Atlas data
Shen, M.; Gao, Y.; Liu, N.; Bhuva, D.; Milton, M.; Henao, J.; Andrews, J.; Yang, E.; Zhan, C.; Liu, N.; Si, S.; Hutchison, W. J.; Shakeel, M. H.; Morgan, M.; Papenfuss, A. T.; Iskander, J.; Polo, J. M.; Mangiola, S.
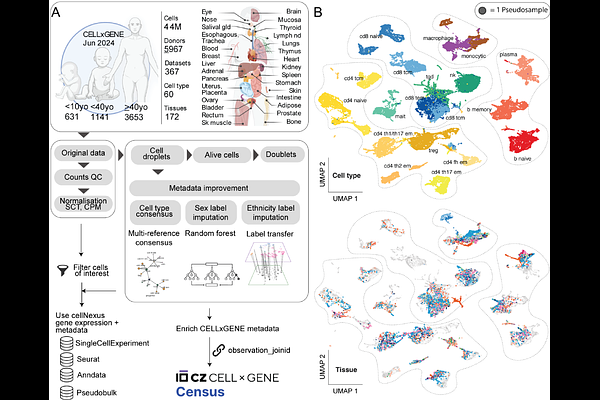
AbstractLarge-scale single-cell atlases such as the Human Cell Atlas have transformed our understanding of human biology. Yet, the lack of a robust framework that standardises quality control, expands cellular annotation, and adds normalisation and analytical layers, limits multi-study analyses and the usefulness of this resource. Here we present cellNexus, a comprehensive tool and resource that converts the Human Cell Atlas collection into analysis-ready data by linking quality control layers, metadata enrichment, expression normalisation, analysis and data aggregation. These enhancements enable robust statistical modelling across studies, exemplified by a multi-tissue map of immune cell communication during ageing, which reveals macrophage-muscle axes as among the most depleted regenerative interactions with age. All harmonised layers, including pseudobulk and cell-cell communication summaries, are accessible via a public web interface and with R and Python APIs. By providing continuous integration with CELLxGENE releases, cellNexus transforms large cell atlas corpora into an accessible, reproducible, interoperable foundation for large-scale biological discovery and the next generation of single-cell foundation models.