The dynamic and heterogeneous structure of the non-canonical inflammasome
The dynamic and heterogeneous structure of the non-canonical inflammasome
Sever, A. I.; Aramini, J. M.; Bonin, J. P.; Zhao, H.; Wang, H.; Rubinstein, J. L.; Schuck, P.; Kay, L. E.
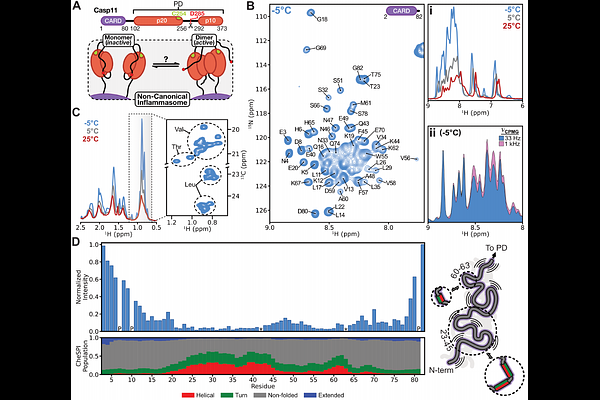
AbstractInflammasomes are high molecular weight complexes that play an integral role in the innate immune system, triggering an inflammatory cascade to protect against cellular stresses such as pathogenic bacteria. Both canonical and non-canonical inflammasomes have been described in the literature and detailed structural studies of many components of the more complex and larger canonical versions have been reported. In contrast, corresponding structures of the non-canonical inflammasome have not emerged even though it consists of only two components: lipopolysaccharide (LPS) from gram-negative bacteria, and one of caspase-4 or caspase-5 in humans or caspase-11 in mice. Here we determine the stoichiometry of the non-canonical inflammasome using size-exclusion chromatography coupled with UV, refractive index, and light-scattering measurements, showing that the non-canonical inflammasome is heterogeneous, comprised of three major complexes with different numbers of LPS and caspase molecules. Solution Nuclear Magnetic Resonance (NMR) spectroscopy studies of the N-terminal Caspase Activation and Recruitment Domain (CARD) of caspase-11, that binds LPS, show that it is largely unstructured in the absence of lipid, with pervasive dynamics on the s-ms timescale. Formation of this complex increases the alpha-helical content of the CARD but the dynamics persist, multiple conformers are formed, and tertiary contacts are transient, consistent with formation of a molten globule. Our NMR results establish that the protease domain of caspase-11 is monomeric in isolation. As proteolysis is linked with dimerization, the protease domains are inactive in this state, but upon formation of the non-canonical inflammasome dimerization occurs, priming the complex for rapid processing of substrates.