Whole-genome 3D architectural screen reveals modulators of brain DNA structure
Whole-genome 3D architectural screen reveals modulators of brain DNA structure
Parasar, B.; Raja Venkatesh, A.; Perera, J.; Sosnick, L.; Moghadami, S.; Seo, Y.; Shi, J.; Chan, L.; Takenawa, S.; Akiyama, T.; Sianto, O.; Uenaka, T.; Hadjipanayis, A.; Wernig, M.; Gitler, A. D.; Tan, L.
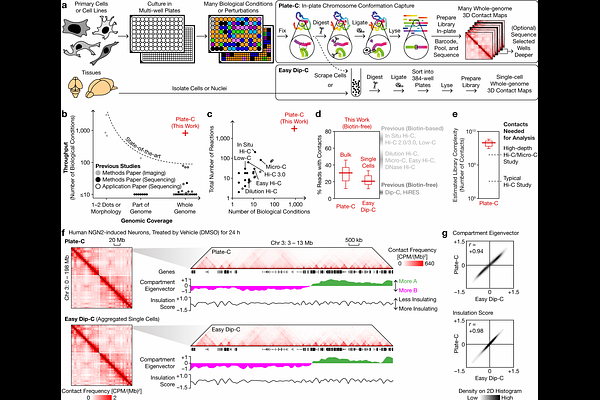
AbstractThree-dimensional (3D) genome architecture is the foundation of gene regulation, and plays a critical role in normal physiology and disease. However, our understanding of its biochemical determinants has long been limited by technology: imaging-based screens only profile a small number of loci, while sequencing-based studies rarely exceed 100 samples or conditions. Here we present "in-plate chromosome conformation capture" (Plate-C), a high-throughput, cost-effective platform that profiles thousands of whole-genome architectures in a day. Plate-C enabled the first chemical screen for whole-genome structural changes--profiling 2,956 samples from 834 conditions across 5 neuronal and glial types, accompanied by 6,081 single cells using "easy diploid chromosome conformation capture" (Easy Dip-C) and 200,893 single-cell transcriptomes. We discovered that diverse, dose/time-dependent, and cell type/species-specific modes of DNA structural changes can be rapidly induced by manipulating epigenetic (HDAC, BET), metabolic (mTOR), proteostatic (UPR), developmental (GSK3/Wnt, Hedgehog), immune (cGAS/STING), and neurotransmission pathways. To validate our finding in vivo, we demonstrated in newborn mice that HDAC inhibition drives brain-wide genome rewiring within hours, highly correlated with changes in vitro and inducing a latent structural and transcriptional state orthogonal to normal differentiation. By enabling massively parallel profiling of whole-genome structures, Plate-C paves the way for systematic discovery of DNA folding principles to better understand and engineer the human genome in 3D.