Resting-state EEG alpha-BOLD coupling spatially follows cortical cell-type and receptor gradients
Resting-state EEG alpha-BOLD coupling spatially follows cortical cell-type and receptor gradients
Jiricek, S.; Chien, V. S. C.; Schmidt, H.; Koudelka, V.; Marecek, R.; Mantini, D.; Hlinka, J.
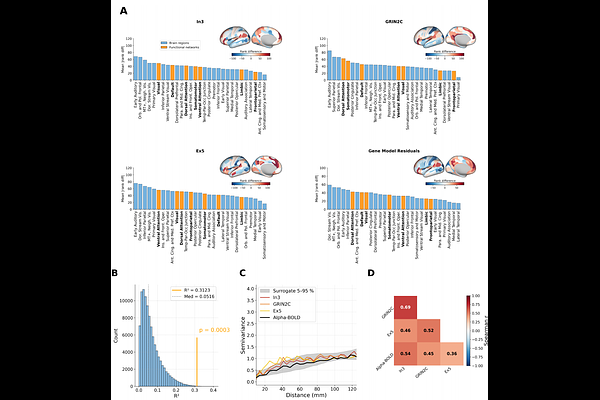
AbstractThe coupling between electroencephalography (EEG) and blood-oxygen-level-dependent (BOLD) signals has been investigated across numerous studies, but its neurobiological underpinnings remain poorly understood. Resting-state EEG alpha-BOLD coupling follows a characteristic spatial pattern, shifting from negative correlations in sensory regions to positive correlations in association cortices. In this study, we examined neurobiological correlates of resting-state alpha-BOLD coupling. We compared the spatial pattern of the alpha-BOLD coupling map to 82 cortical feature maps, including gene expression profiles of different cell types and receptor subunits as well as structural MRI measures. We identified three statistically significant (q < 0.05 FDR-corrected) maps: the layer 6 VIP interneuron marker, excitatory layer-5 marker, and NMDA receptor subunit GRIN2C. The three significant gene maps, combined in a multiple linear regression model, explained R2 = 0.312 of the spatial variance in alpha-BOLD coupling. Analysis of the spatial mismatch between cortical maps and the alpha-BOLD coupling map revealed that the early auditory cortex is the region that consistently diverges from predictions across gene expression and T1/T2 maps. The spatial correspondence between alpha-BOLD coupling and gene expression profiles of specific receptor subunits, neuronal types, and layer-specific populations identifies these as concrete candidates for future computational and experimental studies of alpha-BOLD coupling.