Evolutionary conditioning enables guided generation of functionally diverse enhancers
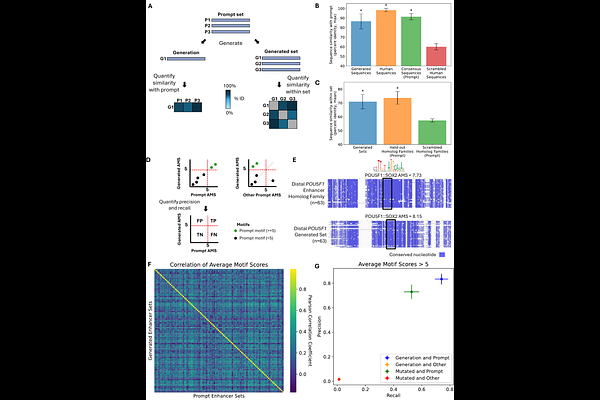
Evolutionary conditioning enables guided generation of functionally diverse enhancers
Duncan, A. G.; Consens, M. E.; Crawford, L.; Mitchell, J. A.; Moses, A. M.; Yang, K. K.; Lu, A. X.
AbstractDeep learning has been instrumental in our understanding of how enhancers encode regulatory information in their DNA sequence and has demonstrated preliminary success with enhancer design. However, the prevailing approach for enhancer design, cell type label conditioning, depends on labeled data from massively parallel reporter assays, which only exists for a handful of cell types. We propose EnhancAR, an autoregressive model trained on sets of unaligned homologous enhancer sequences to learn the function of the enhancer conserved over evolution and generate sequences that resemble real homologs. By training EnhancAR on 1.7 million human enhancer homolog sets spanning 1,888 cell types, EnhancAR generates enhancers for a variety of contexts without being conditioned on a cell type label. We computationally validate that when conditioned on a set of enhancer homologs, EnhancAR generates novel and diverse sequences that preserve the functional properties of the homologs. By prompting EnhancAR with homologs for existing cell type specific enhancers, we design enhancers with similar predicted cell type specificity. We further demonstrate that when trained on length sorted homologs, EnhancAR can design enhancers shorter than the conditioning homologs that preserve the predicted activity. In summary, we find that leveraging evolutionary information in enhancer homologs enables a more flexible and general paradigm for designing enhancers with specific functions.