Genome-wide genealogies reveal deep admixtures forming modern humans
Genome-wide genealogies reveal deep admixtures forming modern humans
Loya, H.; Gupta Hinch, A.; Palamara, P. F.; Speidel, L.; Myers, S. R.
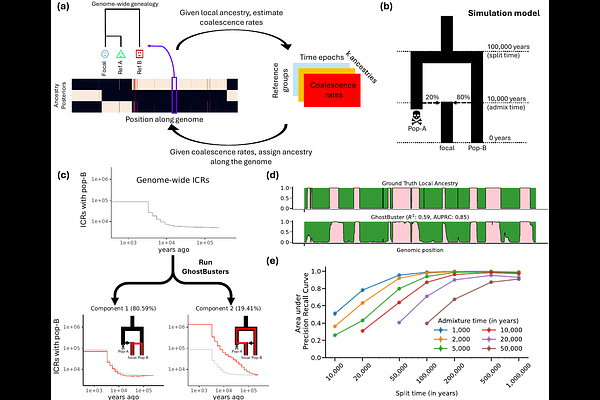
AbstractOver the past decade, genomic modelling has revealed a rich tapestry of admixtures shaping present-day human populations. These have largely focused on the past few thousand years, when ancestral populations are either well characterised by present-day genomic diversity or directly observed through ancient DNA. Genomic modelling and fossil evidence have so far only provided a fragmented picture of the coexistence and mixing of human groups in the deeper past. Here, we propose a new method, GhostBuster, that leverages inferred genome-wide genealogies to detect admixture events of unsampled ghost populations, while simultaneously inferring accurate local ancestry. Local ancestry enables us to identify ancestry-specific genomic signatures that independently corroborate the events. We identify at least three waves of "back-to-Africa" migrations starting ~14,000 years ago. Applying GhostBuster to deeper timescales reveals that modern humans were shaped by repeated episodes of mixture. Around 50,000 years ago, we identify a human lineage that expanded to form present-day non-Africans, while also expanding within Africa, mixing with the other local African group in varying proportions. These ancient groups help explain polygenic score portability differences within Africa, and exhibit differences in population size and recombination landscapes. Extending our analysis further back to between 300,000 and 1 million years ago reveals two deeply diverged ancestral lineages. These lineages evolved profoundly different recombination landscapes, with different PRDM9 alleles (PRDM9-A and C) and recombination hotspots. We demonstrate that both Neanderthals and ancestral modern humans are formed through a mixture of these two lineages, with no evidence of gene flow from the PRDM9-A-carrying group into Denisovans.