Detection of a sequence feature for recursive splicing
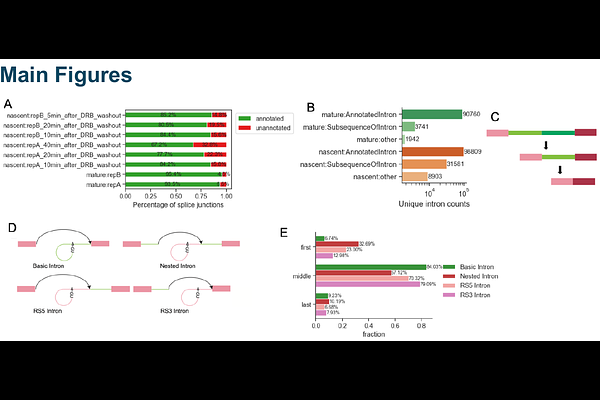
Detection of a sequence feature for recursive splicing
Wang, B.; Yang, K.; Barash, Y.; Choi, P.; Mount, S. M.; Larson, D. R.
AbstractRNA splicing is directed by cis-acting sequence signals interacting with trans-acting factors to remove introns from newly transcribed pre-mRNA, joining exons to generate mature mRNA. Splicing happens far more often than observed exon-exon junctions in mRNA. As one contributing process, the spliceosome progressively removes large introns in small segments by 'recursive splicing', instead of splicing the whole intron at one time. However, the cis-acting sequences associated with recursive splicing have not been identified. Using probabilistic mixture models, we found that recursive splicing occurs more frequently in first introns, which are typically longer, and exhibit a distinct CG-rich sequence feature in the sequences flanking the upstream 5'SS, and depletion of CGs in the downstream polypyrimidine tract. Remarkably, recursive splicing is also more frequent in downstream introns of genes containing first introns with these properties. Mechanistically, these data suggest that early events in RNA synthesis and processing influence the prevalence of recursive splicing for the rest of the transcript. Finally, we developed a sequence-dependent classifier for recursive splicing, which we tested with a novel medium-throughput primer extension assay. In summary, the usage of recursive splicing sites is established at the beginning of RNA synthesis through newly-identified sequence motifs flanking both ends of the first intron.