Refining the Serine Protease Autotransporters of Enterobacteriaceae (SPATE) gene detection in Enteroaggregative Escherichia coli genomes uncovers differential SPATE distribution by phylogeny
Refining the Serine Protease Autotransporters of Enterobacteriaceae (SPATE) gene detection in Enteroaggregative Escherichia coli genomes uncovers differential SPATE distribution by phylogeny
Dada, R. A.; Afolayan, A. O.; Adewuyi, O. A.; Tytler, B. A.; Olayinka, B. O.; Thomson, N. R.; Okeke, I. N.
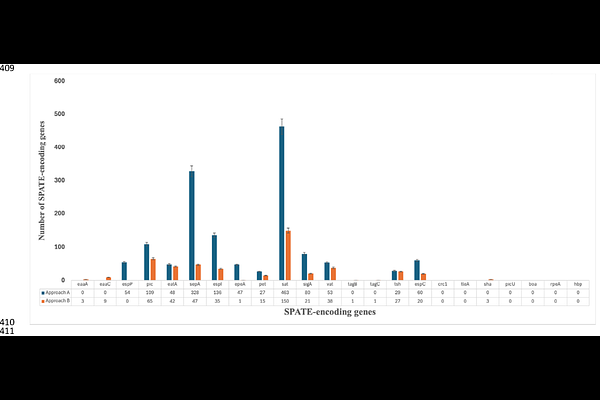
AbstractEnteroaggregative Escherichia coli (EAEC) are a heterogenous pathotype, implicated in acute and persistent diarrhoea especially in developing countries. Serine Protease Autotransporters of Enterobacteriaceae (SPATEs) are Type V Secretory System trypsin-like proteases repeatedly reported from EAEC. This study aimed to determine the prevalence of SPATE encoding-genes among EAEC and their association with diarrhoea. We screened 881 EAEC genomes from four recent epidemiological studies in Nigeria for 23 SPATE-encoding genes, initially using ARIBA and the Virulencefinder database. This initial screening method inflated SPATE gene content, particularly in genomes with multiple SPATEs, due to cross detection of highly similar sequences and other artefacts. We developed and validated refined methodology, which detected 478 of 1,156 original SPATE calls and also identified SPATE miscalls from previous datasets in the literature. The most prevalent SPATE-encoding gene in our EAEC collection was sepA 297(33.71%), closely followed by sat 360 (29.74%). pic, encoding a SPATE with mucinase activity, was found in 65 (7.4%) genomes and associated with diarrhoea (p=0.00004). EAEC strains belonging to E. coli phylogroups A, B1 or C carried, on average, one SPATE gene per genome while >1 was typically detected in phylogroup B2 EAEC. Other EAEC carried few or no SPATE genes. Our study shows that multifunctional genome analysis tools may have to be refined for certain gene families to avoid overestimation. SPATEs are not as prevalent as previously thought but they remain common among EAEC, particularly among phylogroup A, B1, B2 and C, pointing to the possibility that they make lineage-specific contributions to disease.