Dismantling Chromosomal Stasis Across the Eukaryotic Tree of Life
Dismantling Chromosomal Stasis Across the Eukaryotic Tree of Life
Copeland, M.; McConnell, M.; Barboza, A.; Abraham, H. M.; Alfieri, J.; Arackal, S.; Bernard, C. E.; Bryant, K.; Cast, S.; Chien, S.; Clark, E.; Cruz, C. E.; Diaz, A. Y.; Deiterman, O.; Girish, R.; Harper, K.; Hjelmen, C. E.; Thompson, M. J.; Koehl, R.; Koneru, T.; Laird, K.; Lee, Y.; Lopez, V. R.; Murphy, M.; Perez, N.; Schmalz, S.; Sylvester, T.; Blackmon, H.
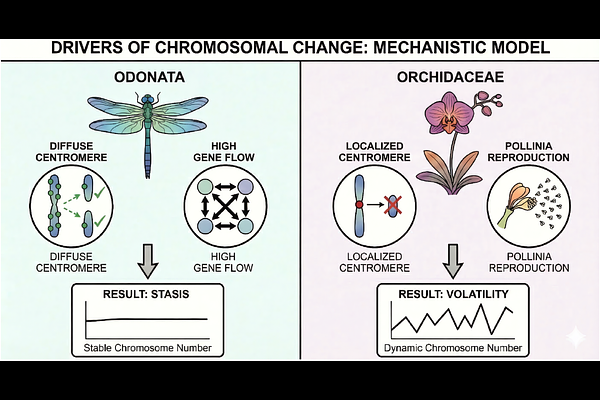
AbstractChromosome number shapes genome organization, recombination, and speciation, yet how fast it evolves across the tree of life has never been measured. We analyzed 63,682 karyotypes across 55 eukaryotic clades and found that dysploidy rates vary by 844-fold, from approximately 0.0008 to 0.7 events per million years. This variation does not follow kingdom boundaries or deep phylogeny; intraclade variance exceeds interclade differences by more than an order of magnitude. Even birds, the textbook example of chromosomal stasis, exceed the global median rate once microchromosome dynamics are resolved. Contrasting the stasis of Odonata with the volatility of Orchidaceae reveals that life history and population structure, rather than deep phylogenetic constraints, govern the tempo of karyotypic change.