CysK2 couples copper sensing to metabolic and redox adaptation in Mycobacterium tuberculosis
CysK2 couples copper sensing to metabolic and redox adaptation in Mycobacterium tuberculosis
Le Mouellic, W.; Levillain, F.; Wu, T.-D.; Larrouy-Maumus, G.; Lagarrigue, F.; Vidal, E.; Rombouts, Y.; Poquet, Y.; Neyrolles, O.
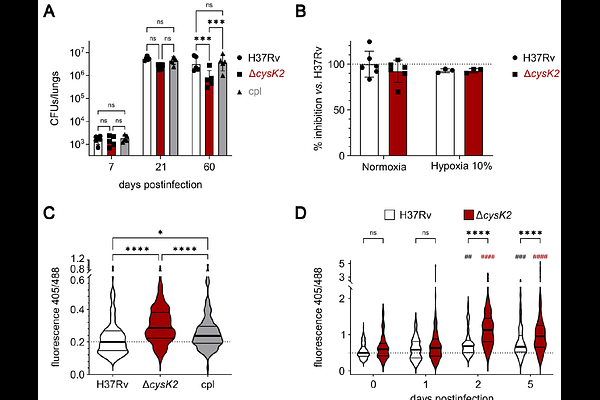
AbstractCopper is increasingly recognized as a host derived cue encountered by Mycobacterium tuberculosis and other microbes during infections, yet the magnitude, intracellular distribution, and physiological consequences of this exposure remain incompletely understood. Here, we combined high resolution imaging, transcriptomic profiling, intracellular reporter assays, isotope tracing, and mouse infection models to define how M. tuberculosis responds to physiologically relevant copper levels during macrophage infection. NanoSIMS analysis showed that copper reaches intracellular bacilli and accumulates in discrete phosphorus rich foci in bacteria. Exposure to physiological copper concentrations in vitro triggered a highly specific transcriptional response dominated by the copper inducible CsoR and RicR regulons. The RicR regulated gene cysK2 gene, encoding the S-sulfocysteine synthase CysK2, was one of the most strongly induced loci. In infected macrophages, cysK2 expression was modulated by extracellular copper availability, host copper transport pathways, and hypoxia. In vitro, an H37Rv cysK2 deficient mutant showed reduced amino acid biosynthesis in response to copper exposure. In vivo the mutant was impaired in long term persistence in mice and displayed a higher oxidation status. Together, these findings identify CysK2 as a copper responsive metabolic effector that couples host derived copper sensing to redox homeostasis and intracellular adaptation in Mycobacterium tuberculosis.